Primary works:
Tutorial: Analytical Protocols for Quantitative Evaluation of [4Fe-4S]-maquette Spectra in support of published results in
Galambas A.; Miller J.; Jones M.; McDaniel E.; Lukes M.; Watts H.; Copié V.; Broderick J.B.; Szilagyi R.K.; Shepard E.M.: Radical S-adenosylmethionine maquette chemistry: Cx3Cx2C Peptide coordinated redox active [4Fe-4S] clusters Journal of Biological Inorganic Chemistry, 2019, 24(6), 793–807 DOI: 10.1007/s00775-019-01708-8.
Step by step instructions for how we model the UV-vis spectra formed by the ligand-field, sulfide, and thiolate charge-transfer bands along with representative worked-out examples are provided at the Zenodo scientific depository.
Szilagyi, R.K.; Hanscam, R.; Shepard, E.M.; McGlynn, S.E.: Natural Selection Based on Coordination Chemistry: Computational Assessment of [4Fe-4S]-Maquettes with Non-coded Amino Acids Interface Focus, 2019, 9(6), Article No.: 20190071 DOI: 10.1098/rsfs.2019.0071 – (Chemobrionics 30/80 themed issue)

- Supporting information is made available about details of every simulations, parameters used for non-natural amino acids at Zenodo repository.
Galambas, A.; Miller, J.; Jones, M.; McDaniel, E.; Lukes, M.; Watts, H.; Copié, V.; Broderick, J.B.; Szilagyi, R.K.; Shepard, E.M.: Radical S-adenosylmethionine Maquette Chemistry: Cx3Cx2C Peptide Coordinated Redox Active [4Fe-4S] Clusters Journal of Biological Inorganic Chemistry, 2019, 24(6), 793–807 (Thematic Issue in celebration of Prof. Joan Broderick’s 2019 ACS Bader Award in bioinorganic and bioorganic Chemistry) DOI: 10.1007/s00775-019-01708-8 – open access

In the foreword of the thematic issue celebrating the scientific achievements of Prof. Joan B. Broderick on the occasion of her receiving the ACS Alfred Bader Award in Bioorganic and Bioinorganic Chemistry at the 2019 ACS Spring National Meeting held in Orlando, FL, Prof. Lawrence Que Jr writes “The the fourth paper of the special issue, Szilagyi and coworkers describe their efforts in comparing the properties of [4Fe-4S]-maquettes made from CxxCxxC fragments bound to 4Fe4S clusters in bacterial ferredoxins with the maquettes derived from CxxxCxxC fragments bound to 4Fe4S clusters in radical SAM enzymes and find them to function equally well in coordinating redox-active [4Fe-4S] clusters.”
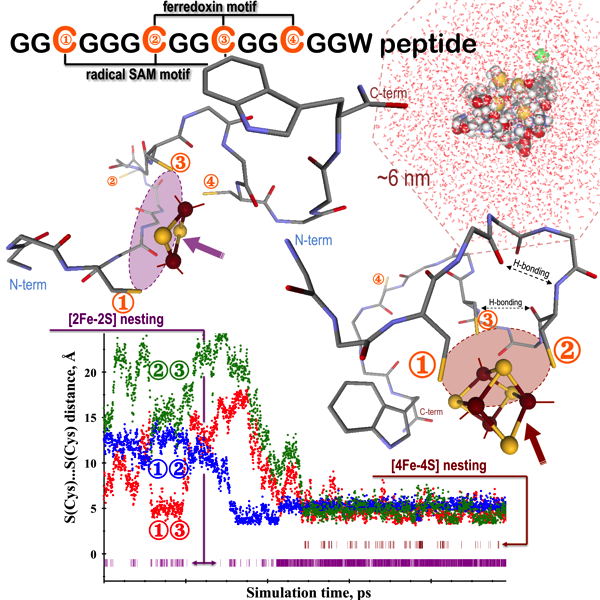
Hanscam, R.; Shepard, E.M.; Broderick, J.B.; Copié, V.; Szilagyi, R.K.: Secondary Structure Analysis of Peptides with Relevance to Iron–Sulfur Cluster Nesting Journal of Computational Chemistry, 2019, 40(2), 515–526 (Special Issue: Memorial Festschrift for Keiji Morokuma) DOI: 10.1002/jcc.25741.
- Related poster presentation from the “Iron-Sulfur Proteins-Biogenesis, Regulation and Function”, 39th Steenbock Symposium.
- An extensive electronic supporting information is available at Zenodo depository including several worked out examples.
Closely related works:
Scott, A.G.; Szilagyi, R.K.; Mulder, D.W.; Ratzloff, M.W.; Byer, A.S.; King, P.W.; Broderick, W.E.; Shepard, E.M.; Broderick; J.B.: Compositional and structural insights into the nature of the H-cluster precursor on HydF Dalton Transactions, 2018, 47(28), 9521–9535 DOI: 10.1039/c8dt01654b
Cited by:
Bortolus, M.; Costantini, P.; Doni, D.; Carbonera, D.: Overview of the Maturation Machinery of the H-Cluster of [FeFe]-Hydrogenases with a Focus on HydFInt. J. Mol. Sci.2018, 19(10), 3118; DOI: 10.3390/ijms19103118
Harris, T.V.; Szilagyi, R.K.: Protein environmental effects on iron-sulfur clusters: A set of rules for constructing computational models for inner and outer coordination spheres Journal of Computational Chemistry, 2016, 37(18), 1681-1696 DOI: 10.1002/jcc.24384
Silver, S.C.; Gardenghi, D.J.; Naik, S.G.; Shepard, E.M.; Huynh, B.H.; Szilagyi, R.K.; Broderick, J.B.: Combined Mössbauer Spectroscopic, Multi-Edge X-ray Absorption Spectroscopic, and Density Functional Theoretical Study of the Radical SAM Enzyme Spore Photoproduct Lyase Journal of Biological Inorganic Chemistry, 2014, 19(3), 465-483. DOI: 10.1007/s00775-014-1104-y